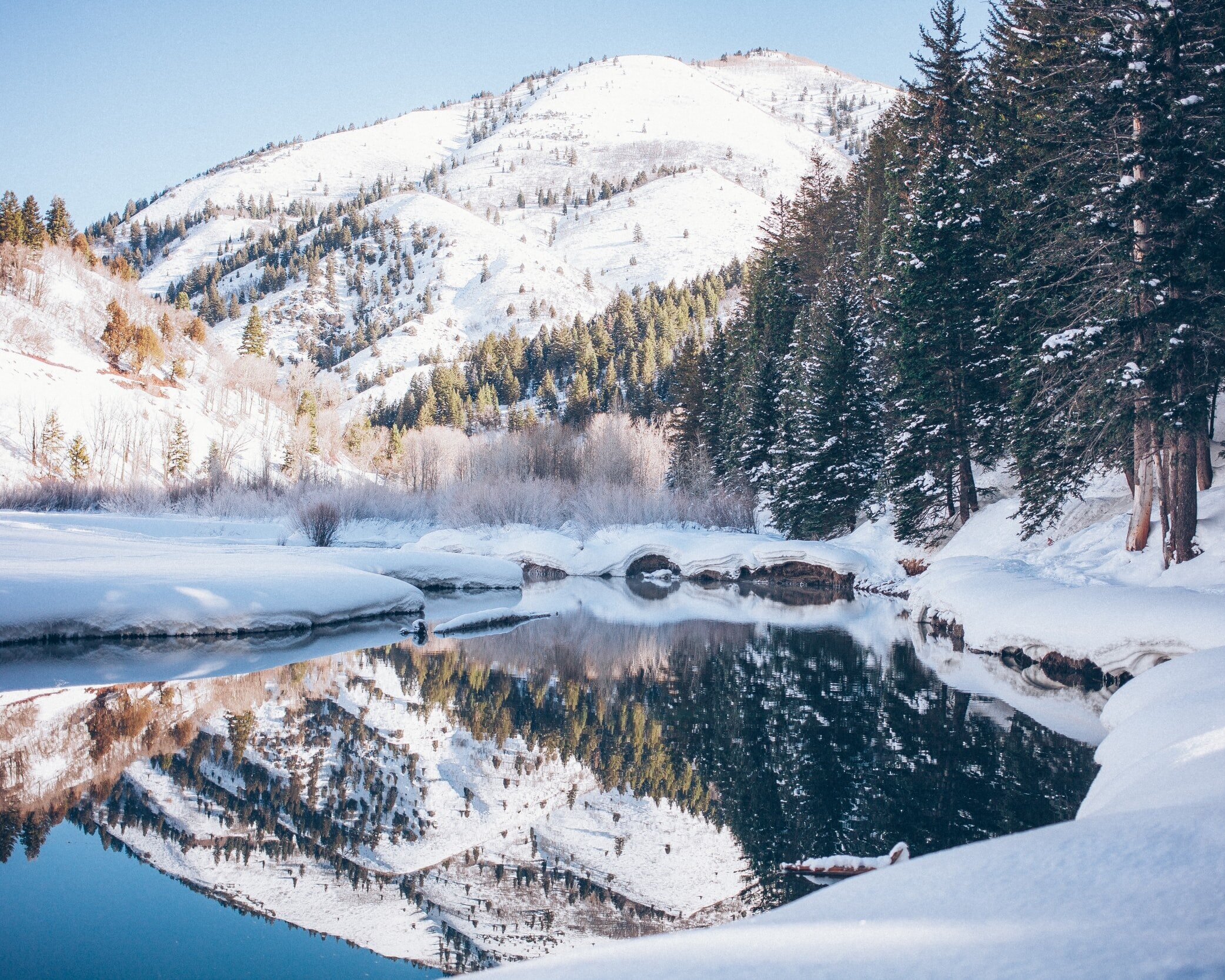
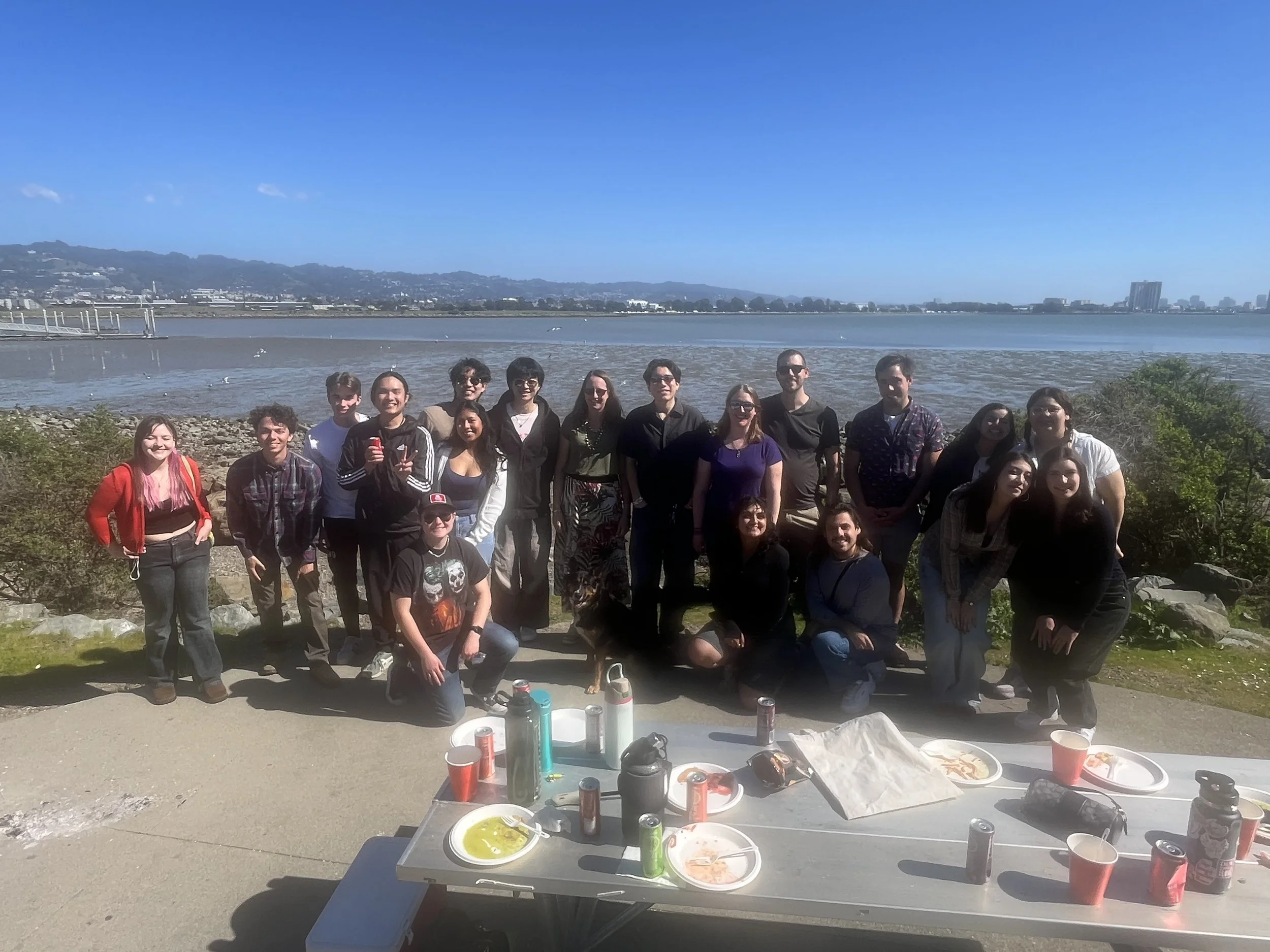
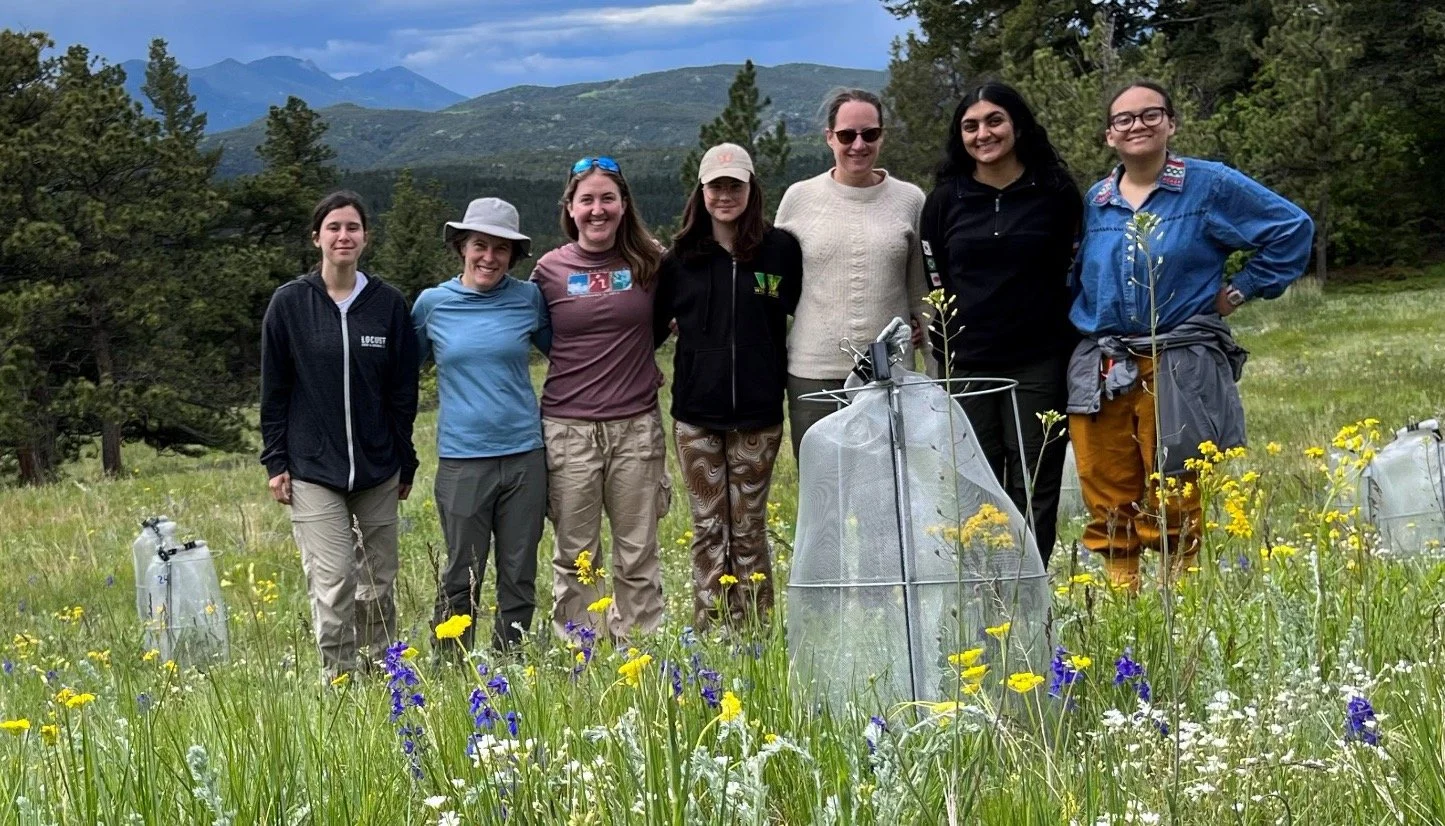
Here in the Williams Lab, we study fundamental questions about change.
We explore the evolutionary impacts of seasonality on performance and fitness using a variety of insect species. Our goal is to support and nurture the diversity of life by studying the natural world.
Our study systems
Latest News in Ecophysiology
About our lab
Diversity is our strength
We are committed to transforming academia to welcome and support all scientists, broadening the picture of what a scientist looks like so that we can benefit from the full pool of talent that a diverse community brings. We work actively towards a more equitable, just and inclusive scientific community.

Join the conversation!
-
We have lab beetle t-shirts or hoodies available! If you want to buy one ($20 t-shirts, $40 hoodies) hit me up! Tha… https://t.co/I2d7aUoIRq
-
RT @JessicaHellmann: Big announcement and big opportunity for emerging leaders to be part of @UMNIonE. https://t.co/l5SSjfSuiN
-
RT @bkoskella: [Please retweet!] Postbac research opportunity: Bay Area RaMP in Microbiome Sciences (https://t.co/KB3hhuKlZj) offe… https://t.co/xoYiC4KwGp